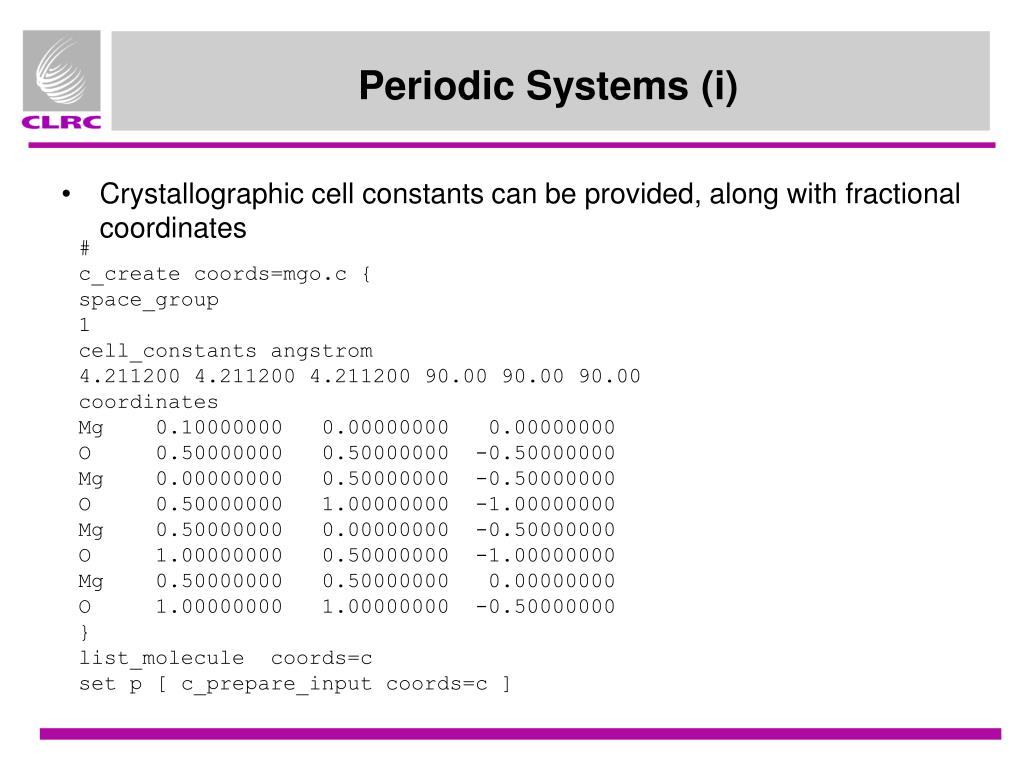
  Post memes/jokes in /r/chemistrymemes and /r/chemistryjokes.  Any such posts will be deleted.Īsk education and jobs questions in the current weekly topic. If you're looking for a more concentrated, advanced discussion of chemistry topics among professionals and grad students, check out /r/Chempros.Ä«efore asking "What chemical is this?" see this chart.  Click here for the OSHA chemical data site and here for a multicompany MSDS aggregate search. If you spill/injure yourself contact medical professionals and read the MSDS, do not post to this reddit. I need to specify certain coordinates in the matrix, therefore, the automatic built-in function from gaussview isn't of much help. Yes links to blogs, images, videos, comics, and infographics are okay especially if they are on your personal website. Z-Matrix Generator for Gaussian Ask Question Asked 6 years, 1 month ago Modified 9 months ago Viewed 7k times 4 I am looking for a way to convert XYZ coordinates into z-matrix for Gaussian input files. In this node, the number of ones in each column and the number of. No physorg, sciencedaily, or other press release aggregator spam! Generates a regular parity check matrix that is used by MT LDPC Encoder for LDPC encoding. If a caption or explanation is included this helps, but please use your discretion.Ä«efore asking about chemical drawing/illustration programs, look at your school's IT/software website and see if they provide an institutional license of ChemDraw (hint: if they have a chemistry department, they will) Likewise, simple pictures of uninteresting and garden variety chemistry-related things are not appreciated. The MULTICI generates a matrix file that describes the averaged read density in specified BED site. For now we will generate actual and predicted. MULTICI: Generate matrix of averaged read density. No memes, rage comics, image macros, reaction gifs, or other "zero-content" material. Confusion matrixes can be created by predictions made from a logistic regression. Robust and exhaustive processes to generate chemical structures have been described previously in different contexts 1, 2, 3, 4, 5, 6, 7, 8, 9. However, academic discussions on pharmaceutical chemistry and the science of explosives are permitted. Introduction Initial structures (Z-matrix or Cartesian coordinate) are important starting points for the in silico investigation of chemical species. This gives you the possibility to check and compute manually (or, calculator) the distances within reasonable effort to check if the algorithm works well. (END ZEND) The input module recognizes three possible spellings of this directive name. Start with a smaller molecule in first place. But if your constraints do not consider the distribution, this could do as well (and this will give you a matrix from a single run).Rules: Violating a rule will result in a ban.Īsk homework, exam, lab, and other undergraduate-level questions at ChemicalForums otherwise it will be deleted.Äiscussions on illicit drug synthesis, bomb making, and other illegal activities are not allowed and will lead to a ban. The general form of the compound ZMATRIX directive is as follows: ZMATRIX ZMT ZMAT .Note that as pointed out, this will have a different distribution as the rejection method. For practical reasons I padded the matrix with a zero line and column at the beginning to avoid indexing problems with k1-1 and k2-1. It's a bit tricky because for each element you check the row-sum and column-sum until that point, and use the larger of the two for generating a new element (in order to stay below a sum of 1 in both directions). XYZ coordinates have a redundancy, as of the 3N coordinates, only 3N-6 are unique. My first method work with one class and complex (self.real,self.imag) : Operator overloading +- is ok. 218 21K views 6 years ago Computational Chemistry Short lecture on the z-matrix format for molecular coordinates. Matrix(k1,k2)=rand()*(1-max(sum(matrix(k1,1:k2-1)),sum(matrix(1:k1-1,k2)))) How create complex number matrix in Python Ask Question Asked 2 years, 3 months ago Modified 2 years, 3 months ago Viewed 46 times -1 I am trying to create complex number matrix class using Python. For Targeted Gene Expression samples, non-targeted genes are removed from the filtered matrix. Matrix = zeros(n+1) %dummy line in first row/column Contains only detected cell-associated barcodes. For this you always generate a new element between 0 and the remainder until 1: n = 3 of existing random number generators, a matrix A whose elements have the same joint distribution as the elements of A. So another, but more tedious approach is to generate each element such that the sum can only be 1 in each direction. The rejection method will surely give you a uniform solution, but it might take a long time to generate a good matrix, especially if your matrix is large. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
